Lighting the LAMP to Reveal Mystery of Lysosomes
Tokyo, Japan – A cell is composed of numerous organelles, each with a unique role that helps contribute to its overall functionality. The lysosome is an organelle that contains digestive enzymes and functions as a molecular garbage disposal and recycling center. Since the role of lysosome is crucial to maintain the cellular homeostasis, the lysosomal dysfunction causes neurodegenerative and metabolic diseases, cancer, as well as lysosomal storage disorders.
In a new article published in Autophagy, researchers at Tokyo Medical and Dental University (TMDU) performed a novel type of structural analysis to demonstrate how a certain molecular interaction is crucial for one lysosomal membrane protein to perform effectively.
LAMP1 (lysosomal-associated membrane protein 1) and LAMP2 the most abundant protein components of lysosome membranes. Both LAMP1 and LAMP2 are composed of a large lumenal domain, a transmembrane domain, and a short C-terminal cytoplasmic tail (Fig. 1). The lumenal domains of LAMPs are composed of two domains (N-domain and C-domain, which are membrane-distal and -proximal, respectively). Each domain has the unique β-prims fold structure (a triangular prism). On the other hand, genetic experiments have shown that mice embryos without both LAMP-1 and 2 die a little more than two weeks after fertilization. Mice without LAMP-1 are born and can thrive, while those without LAMP-2 often die a few weeks after birth, suggesting that LAMP-2 is more important for lysosome activity. The research of the TMDU group further investigates this protein’s biological role.
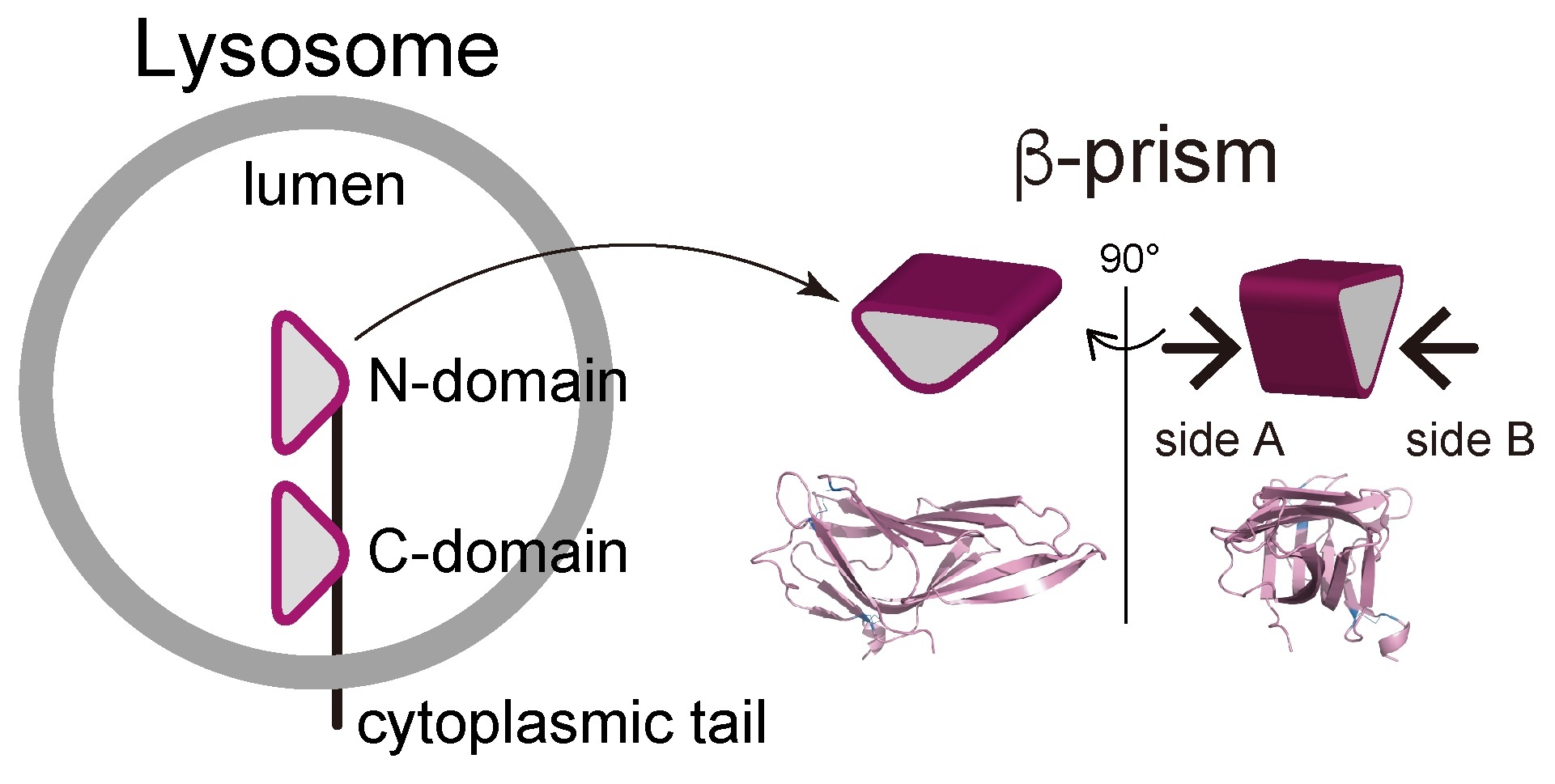
Figure.1 The two-domain architecture of LAMP1 and LAMP2
Domain architecture and orientation of LAMP1 and LAMP2 in the lysosome lumenal membrane(left). Schematic β-prism shapes and the structures of the C-domain of LAMP1(5GV0) are shown in two perpendicular views(right).We defined the surface of the N-terminal(N) side as SideA, and that on the C-terminal(C)side as Side B.
The researchers demonstrated LAMP2 molecules assemble by facing each other with one side the β-prism (defined as side A, as shown in Fig. 1) of the C-domain (Fig. 2). The N-domain truncation permitted the nonspecific involvement of both sides of the β-prims (side A and side B). In combination with some biochemical studies, “we believe that the homophilic interaction we demonstrated is crucial for function of LAMP-2, via ensuring a proper arrangement of the cytoplasmic tails, which is crucial for the function of LAMP2, on the lysosome membrane,” says Miki Hara-Yokoyama, senior author.
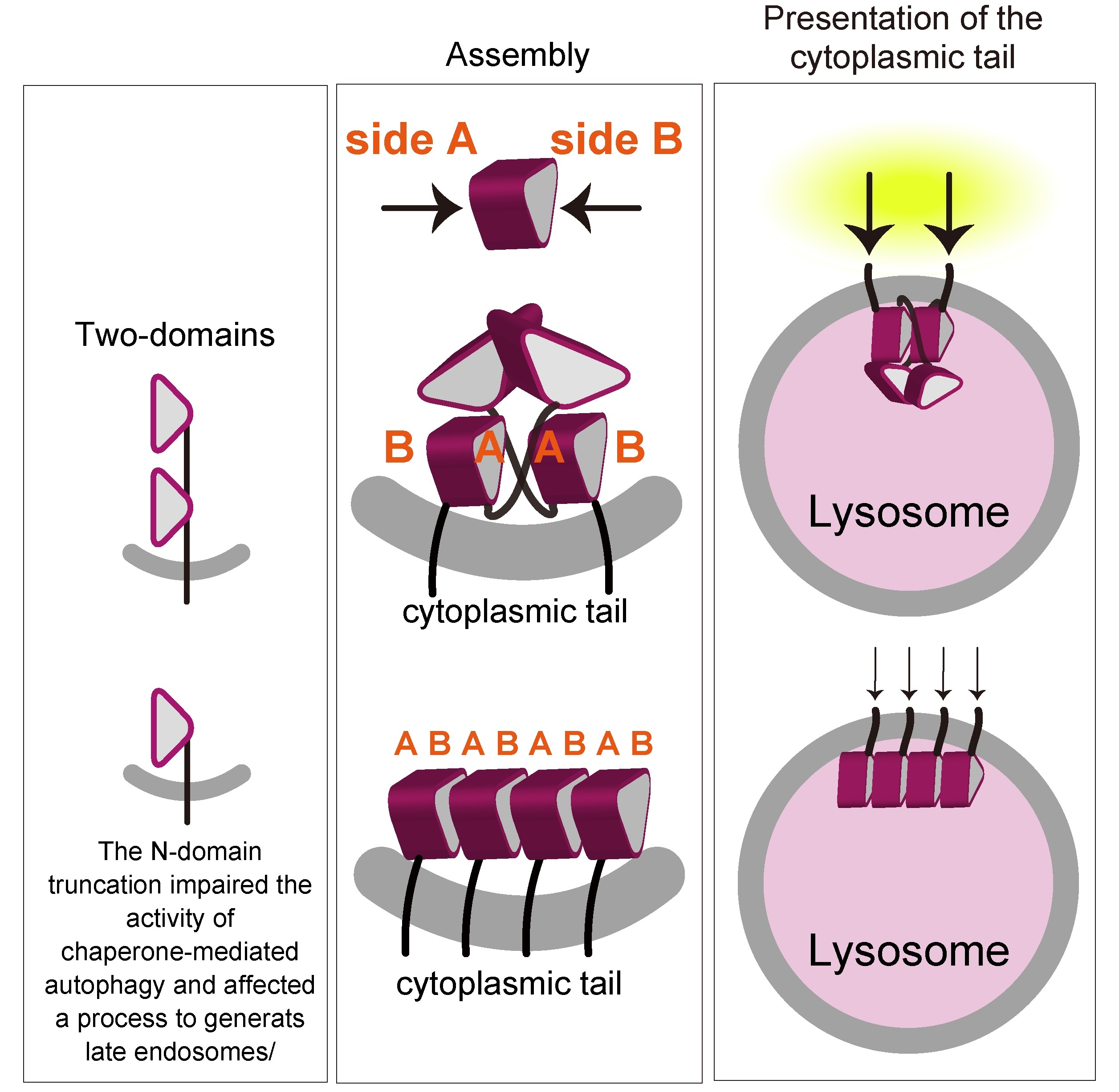
Figure.2 Effect of the truncation of one β-prism domain
Schematic representation of the interaction of the full-length LAMP2A molecules and the N-domain truncated LAMP2A molecules on the lumenal membrane
Summary
Journal Article
TITLE:Direct homophilic interaction of LAMP2A with the two-domain architecture revealed by site-directed photo-crosslinks and steric hindrances in mammalian cells
DOI:https://doi.org/10.1080/15548627.2021.1911017
Correspondence to
Miki Hara-Yokoyama,Associate Professor
Department of Biochemistry,
Graduate School of Medical and Dental Sciences,
Tokyo Medical and Dental University(TMDU)
E-mail:yokoyama.bch(at)tmd.ac.jp
*Please change (at) in e-mail addresses to @ on sending your e-mail to contact personnels.

