Seeing Lipids More Deeply with Mass Spectrometry
In the past, due to various reasons related to their complexity and their low intracellular concentrations, the profiling of these lipids and the linking of a specific acyl variant to biological change has been difficult. However, a new system called PRMC-MS (Phosphoinositide Regioisomer Measurement by Chiral column chromatography and Mass Spectrometry) has now enabled the characterization of the dynamics of phosphoinositide acyl variants both in intracellular and extracellular environments.
Previous methods of measuring and profiling phosphoinositides have produced results that cannot be easily applied to clinical or pathological samples from experimental animals. Even newer methods involving the use of mass-spectrometry which have made advances in some areas still reflect the problem of how to simultaneously quantify the acyl variants of individual regioisomers in biological samples.
The PRMC-MS method now solves this problem and points the way to an understanding of how these lipids influence cell functions. Using PRMC-MS, it is now possible to simultaneously measure all eight classes of phosphoinositides in a single sample. The highly sensitive nature of PRMC-MS allows for the detection of tiny but important changes in intracellular phosphoinositide levels, yielding data that shows that it can be applied to blood samples to track phosphoinositide signatures potentially related to disease states.
PRMC-MS enables the comprehensive analysis of phosphoinositide acyl variants in various types of biological samples, including surgical specimens, which can be used to throw a light on previously unrecognized disturbances of phosphoinositide fatty acyl profiles in cancerous tissue and to monitor their extracellular mobilization. Further study of the differing acyl variants and their conferring of protein binding properties could possibly also reveal how they activate a signaling pathway that favors cancer cell growth and survival and emerge as a target for cancer therapy. Thus, PRMC-MS may well illuminate the role played by phosphoinositides in the pathogenesis of cancers and inflammatory diseases.
In addition, the use of PRMC-MS in the evaluation of phosphoinositide signatures at the acyl variant level in tissue and liquid biopsies may reveal biomarkers suitable for a wide variety of clinical applications.
In the future, applications such as the above may greatly facilitate drug development strategies based on the devising of a therapeutic agent that pinpoints a specific pathogenic phosphoinositide acyl variant, and thus open the way for much more accurate therapeutic methods and cures for patients suffering from a range of diseases that have proven difficult in the past.
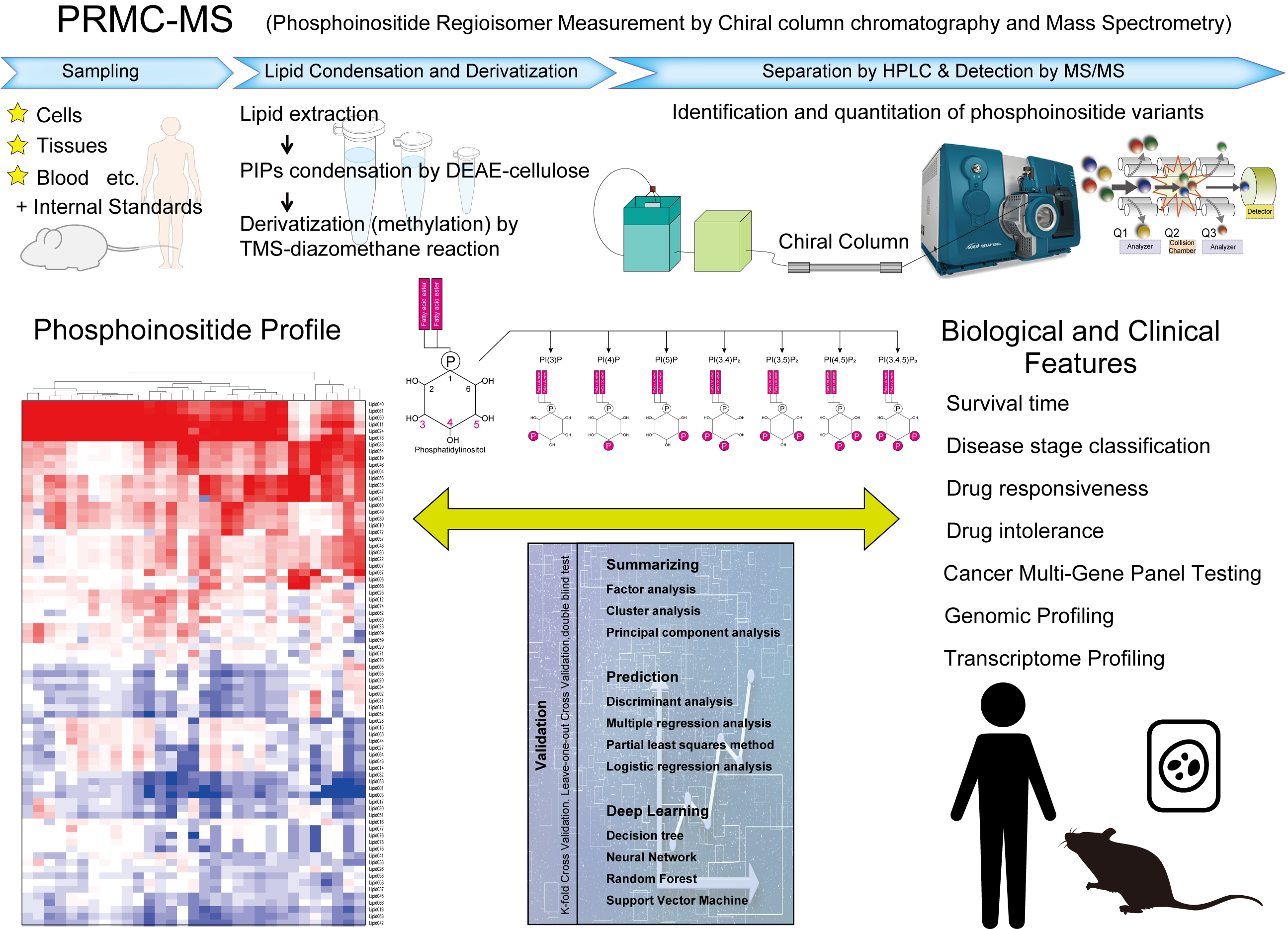
Phospholipid analysis for disease diagnosis and therapy
PRMC-MS enables in-depth profiling of phosphoinositides and paves the way for medical applications of bioactive phospholipids.
Summary
PRMC-MS allows enhanced profiling of phosphoinositide acyl variants both in intracellular and extracellular environments.
Journal Article
TITLE:A mass spectrometric method for in-depth profiling of phosphoinositide regioisomers and their disease-associated regulation
DOI:https://doi.org/10.1038/s41467-021-27648-z

