How Cells Decide the Way They Want to Recycle Their Content
Researchers from Tokyo Medical and Dental University (TMDU) identify a novel phosphorylation site of the protein Ulk1 to be an essential regulating factor for alternative autophagy, but not canonical autophagy
Tokyo, Japan – Autophagy is a housekeeping process through which cells remove dysfunctional contents to balance energy sources during times of stress. Now, researchers from Tokyo Medical and Dental University (TMDU) identified a novel molecular mechanism by which a type of autophagy, called alternative autophagy, is activated. In a new study published in Nature Communications, they showed how a specific phosphorylation site of the protein Unc51-like kinase 1 (Ulk1) is essential for the cell to go down the alternative autophagy path.
As living structures, cells ensure homeostasis by carrying out specific processes by which they build and degrade their contents. Particularly at times of stress, for example during exposure to toxins, autophagy helps to ensure an orderly turnover process by which cells can recycle their contents to survive. Interestingly, the process of autophagy can take place by several distinct molecular mechanisms, two of which are canonical and alternative autophagy. While the protein Ulk1 is known to initiate both types of autophagy, the mechanism by which Ulk1 differentially regulates them has remained unclear.
“Autophagy is a very elaborate process by which cells recycle their contents,” says the corresponding author of the study Shigeomi Shimizu. “The goal of our study was to understand how Ulk1 that has control over two types of autophagy, differentially regulates them.”
To achieve their goal, the researchers used mouse embryonic fibroblasts (MEFs) deficient in the protein Atg5 to turn off canonical autophagy. By exposing them to etoposide, a DNA-damaging reagent, they then induced alternative autophagy. Using mass spectrometry, the researchers found that Ulk1 carried an additional phosphoryl group at its amino acid serine in position 746 (Ser746; p-Ulk1746), also called phosphorylation, when exposed to etoposide but not when left untreated. By developing a new antibody against p-Ulk1746, the researchers then showed that the protein localized to the Golgi complex within the cells. The Golgi complex is an organelle participating in many cellular processes, including alternative autophagy.
“While these were already exciting findings, our goal was to understand whether the specific phosphorylation of Ulk1 at the serine 746 site is required for alternative autophagy and which kinase is responsible for this phosphorylation step,” says lead and the corresponding author of the study Satoru Torii.
To analyze the causal relationship between Ulk1 Ser746 phosphorylation and alternative autophagy, the researchers used a fluorescent tandem protein consisting of red fluorescent protein (RFP) and green fluorescent protein (GFP). Because GFP does not fluoresce within acidic environments, the tandem protein made autolysosomes, cellular compartments that are created during autophagy, become red. While the red fluorescence appeared after etoposide treatment, it was not generated in cells producing Ulk1 nonphosphorylated mutant, indicating that p-Ulk1746 is required for alternative autophagy. Next, the researchers demonstrated that receptor-interacting protein kinase 3 (RIPK3), a protein that phosphorylates other proteins involved in necroptosis, is responsible for the generation of p-Ulk1746 by showing that p-Ulk1746 and alternative autophagy occurred in normal cells but not in cells deficient in RIPK3. Intriguingly in MEFs that expressed Atg5, canonical autophagy was not affected by RIPK3-deficiency, indicating that p-Ulk1746 is not involved in canonical autophagy.
Tokyo, Japan – Autophagy is a housekeeping process through which cells remove dysfunctional contents to balance energy sources during times of stress. Now, researchers from Tokyo Medical and Dental University (TMDU) identified a novel molecular mechanism by which a type of autophagy, called alternative autophagy, is activated. In a new study published in Nature Communications, they showed how a specific phosphorylation site of the protein Unc51-like kinase 1 (Ulk1) is essential for the cell to go down the alternative autophagy path.
As living structures, cells ensure homeostasis by carrying out specific processes by which they build and degrade their contents. Particularly at times of stress, for example during exposure to toxins, autophagy helps to ensure an orderly turnover process by which cells can recycle their contents to survive. Interestingly, the process of autophagy can take place by several distinct molecular mechanisms, two of which are canonical and alternative autophagy. While the protein Ulk1 is known to initiate both types of autophagy, the mechanism by which Ulk1 differentially regulates them has remained unclear.
“Autophagy is a very elaborate process by which cells recycle their contents,” says the corresponding author of the study Shigeomi Shimizu. “The goal of our study was to understand how Ulk1 that has control over two types of autophagy, differentially regulates them.”
To achieve their goal, the researchers used mouse embryonic fibroblasts (MEFs) deficient in the protein Atg5 to turn off canonical autophagy. By exposing them to etoposide, a DNA-damaging reagent, they then induced alternative autophagy. Using mass spectrometry, the researchers found that Ulk1 carried an additional phosphoryl group at its amino acid serine in position 746 (Ser746; p-Ulk1746), also called phosphorylation, when exposed to etoposide but not when left untreated. By developing a new antibody against p-Ulk1746, the researchers then showed that the protein localized to the Golgi complex within the cells. The Golgi complex is an organelle participating in many cellular processes, including alternative autophagy.
“While these were already exciting findings, our goal was to understand whether the specific phosphorylation of Ulk1 at the serine 746 site is required for alternative autophagy and which kinase is responsible for this phosphorylation step,” says lead and the corresponding author of the study Satoru Torii.
To analyze the causal relationship between Ulk1 Ser746 phosphorylation and alternative autophagy, the researchers used a fluorescent tandem protein consisting of red fluorescent protein (RFP) and green fluorescent protein (GFP). Because GFP does not fluoresce within acidic environments, the tandem protein made autolysosomes, cellular compartments that are created during autophagy, become red. While the red fluorescence appeared after etoposide treatment, it was not generated in cells producing Ulk1 nonphosphorylated mutant, indicating that p-Ulk1746 is required for alternative autophagy. Next, the researchers demonstrated that receptor-interacting protein kinase 3 (RIPK3), a protein that phosphorylates other proteins involved in necroptosis, is responsible for the generation of p-Ulk1746 by showing that p-Ulk1746 and alternative autophagy occurred in normal cells but not in cells deficient in RIPK3. Intriguingly in MEFs that expressed Atg5, canonical autophagy was not affected by RIPK3-deficiency, indicating that p-Ulk1746 is not involved in canonical autophagy.
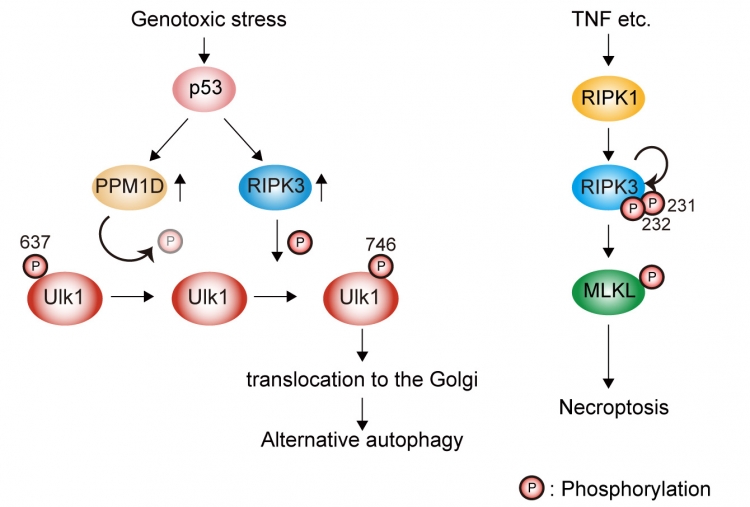
Figure:Elucidation of regulatory mechanism of Ulk1 in alternative autophagy.
Genotoxic stress induces Ulk1 dephosphorylation at Ser637 in a p53/PPM1D-dependent manner. The dephosphorylated Ulk1 is then phosphorylated at Ser746 by a necroptosis initiator, RIPK3, and translocates to the Golgi, which results in alternative autophagy (left). TNF signaling induces necroptosis but not alternative autophagy (right).
“These are striking results that shed new light on how cells regulate the complex process of autophagy,” says Shimizu. “We hope that our findings will be helpful in understanding the role of alternative autophagy in normal biology and disease.”
The article, “Identification of a phosphorylation site on Ulk1 required for genotoxic stress-induced alternative autophagy,” was published in Nature Communications at DOI: 10.1038/s41467-020-15577-2
The article, “Identification of a phosphorylation site on Ulk1 required for genotoxic stress-induced alternative autophagy,” was published in Nature Communications at DOI: 10.1038/s41467-020-15577-2
Summary
Researchers from Tokyo Medical and Dental University (TMDU) identified a new phosphorylation site of Ulk1 as a novel regulating mechanism of alternative autophagy. By inducing DNA damage in MEFs using etoposide, the researchers found that Ulk1 became phosphorylated at the serine 746 site by RIPK3 in alternative autophagy, but not canonical autophagy (see Figure). These findings could help to understand the role of alternative autophagy in normal biology and disease.
Correspondence to
TORII Satoru, Ph.D., Junior Associate Professor
SHIMIZU Shigeomi, M.D., Ph.D., Professor
Pathological Cell Biology,
Medical Research Institute,
Tokyo Medical and Dental University(TMDU)
E-mail:toripcb(at)tmd.ac.jp
shimizu.pcb(at)mri.tmd.ac.jp
*Please change (at) in e-mail addresses to @ on sending your e-mail to contact personnels.
SHIMIZU Shigeomi, M.D., Ph.D., Professor
Pathological Cell Biology,
Medical Research Institute,
Tokyo Medical and Dental University(TMDU)
E-mail:toripcb(at)tmd.ac.jp
shimizu.pcb(at)mri.tmd.ac.jp
*Please change (at) in e-mail addresses to @ on sending your e-mail to contact personnels.

