Know your enemy to know your strategy: Identification of genomic signatures and biochemical pathways in precision medicine
PDF Download
- Aiming at precision medicine

Johji Inazawa
Professor of Molecular Cytogenetics at TMDU
Profile
Dr. Inazawa graduated from Kyoto Prefectural University of Medicine where he received his MD and PhD. He pursued postdoctoral research at Kyoto Prefectural University from 1982 to 1996, when he became Associate Professor at the University of Tokyo. He joined TMDU as Professor of Molecular Cytogenetics at Medical Research Institute in 1998,and assumed the position of Director of the Bioresource Research Center in 2012.

Toshihiro Tanaka
Professor of Human Genetics and Disease Diversity at TMDU
Profile
Dr. Tanaka graduated from the University of Tokyo, School of Medicine where he received his MD and PhD. He conducted research as Assistant Professor at the University of Tokyo from 1997 to 1999, then moved to RIKEN in 2000, and took the post of Deputy Director of RIKEN Center for Genomic Medicine in 2009. Since 2013, he has been Professor of Research Division at the Bioresource Research Center of TMDU and Professor of Human Genetics and Disease Diversity at Graduate School of TMDU.
A:The TMDU Bioresource Research Center aims to discover new causes for diseases and develop precision medicine—a key concept in recent years. Precision medicine is disease treatment and prevention that takes into account individual variability in genes, environment, and lifestyle. This enables us to more accurately predict which treatment and prevention strategies for a particular disease will work in which groups of people. Research has already revealed many of the molecular mechanisms that lead to diseases, with some having their own unique genomic signatures and biochemical pathways. While the concept of precision medicine is not new—blood typing, for instance, has been used to guide blood transfusions for more than a century—the prospect of applying this concept broadly has been dramatically improved by the recent development of large-scale biologic databases, powerful methods for characterizing patients, and computational tools for analyzing large sets of data. At the Center, we have advanced facilities to collect and maintain high-quality data, as well as tissue samples, from patients at TMDU hospitals.
A: The protein NRF2 plays a key role in cellular antioxidant defenses and maintains redox homeostasis. A substantial body of literature demonstrates the enhancement of NRF2 function as a promising antioxidant strategy. In human cancers, aberrantly stabilized NRF2, either by mutation of the NRF2 or KEAP1 gene, plays a vital role in chemoresistance and tumor cell growth, suggesting that targeted inhibition of NRF2 is a potential therapy for NRF2-stabilized tumors.
MicroRNAs (miRNA), which are small RNAs that are not directing the production of a peptide sequence, can negatively regulate gene expression by interfering with the translation and/or the stability of target transcripts. Cancer cells are thought to upregulate cytoprotective processes (protection of cells by chemical compounds) for their survival. Previously, we reported that overexpression of the miRNA, miR-634, activates enhanced chemotherapy-induced cytotoxicity in a model of esophageal cancer, where resistance to chemotherapy remains clinically problematic (Mol. Cancer Res. 2014, Cancer Res. 2015). More recently, we found that inhibition of another miRNA, miR-432-3p, results in increased sensitivity of esophageal cancer cells to chemotherapy drugs, including cisplatin.
Taken together, our findings provide novel insights for regulation of the NRF2 pathway and for miRNAs as targets for overcoming chemoresistance in patients with cancer.
A: To further define the genetic basis of atrial fibrillation, we conducted a meta-analysis of 22,346 individuals with atrial fibrillation and 132,086 referents as international collaborative efforts. We successfully identified 12 new genetic loci, implicating genes involved in cardiac electrical and structural remodeling. We focused on atrial fibrillation because it is a common cardiac arrhythmia condition that can cause serious complications such as stroke, heart failure, dementia, and death. The lifetime risk of atrial fibrillation is one in four, with over 33 million individuals estimated to be affected worldwide today.
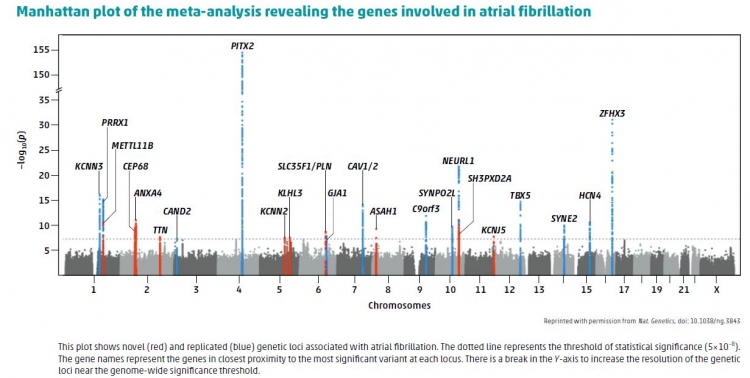
A: Our results nearly double the number of known genetic loci implicated in atrial fibrillation,providing novel insights into the molecular basis of atrial fibrillation. We have identified a series of new atrial fibrillation associated loci variants, which lie proximal to genes involved in atrial electrical and mechanical function. This is valuable information for future downstream research to establish the mechanistic links between identified genetic loci and atrial fibrillation pathogenesis. Most importantly, our findings may help in the discovery of new therapeutic targets for the treatment of atrial fibrillation.
Journal Information
Nat. Genetics, doi: 10.1038/ng.3843

